What is Acute Myeloid Leukemia (AML)?
Acute Myeloid Leukemia (AML) is a highly aggressive and challenging hematologic disease characterized by the rapid proliferation of abnormal myeloid blasts in the bone marrow and peripheral blood. AML is notoriously difficult to eradicate because leukemic stem cells can hijack receptors like CXCR4 to anchor themselves within the bone marrow, resulting in resistance to chemotherapy. These residual leukemic blasts undergo clonal evolution, leading to high relapse rates even after patients achieve an initial clinical remission.
The role CXCR4 and CXCL12 play in AML
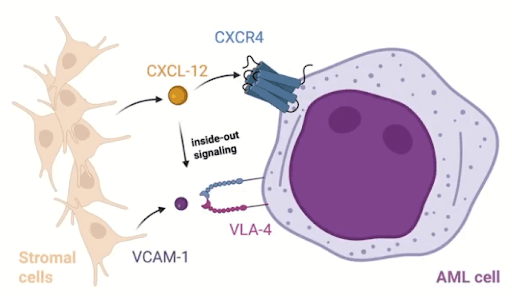
The biological axis of CXCR4 and CXCL12 is foundational to both normal hematopoiesis and AML pathogenesis. CXCR4 is a G-protein coupled receptor that natively binds to its primary ligand, CXCL12, which is also commonly referred to as Stromal Cell-Derived Factor 1 or SDF-1. Within the normal bone marrow microenvironment, SDF-1 is primarily secreted by bone marrow stromal cells and other niche-supporting cells. When the CXCR4 receptor is activated by CXCL12, it initiates a cascade of downstream intracellular signaling pathways. These pathways include the PI3K/AKT pathway, the MAPK signaling cascade, and the JAK/STAT signaling pathway. The collective activation of these pathways promotes cellular survival, proliferation, and directed migration.
In healthy physiological states, this axis regulates stem cell homing and retention within the protective bone marrow. However, in the context of Acute Myeloid Leukemia, leukemic blasts actively hijack this physiological mechanism. Leukemic cells that overexpress CXCR4 remain tightly anchored within the protective microenvironment of the bone marrow. Within this localization, the AML blasts have reduced susceptibility to the cytotoxic effects of chemotherapy and also evade susceptibility to immune-mediated clearance. This localized protection contributes directly to clonal persistence and disease relapse.
The mechanistic context involves activation of integrin-mediated signaling pathways, specifically VLA-4 and VLA-5. The activation of these integrins ensures enhanced adhesion of the leukemic cells to Vascular Cell Adhesion Molecule 1 (VCAM-1) and fibronectin located within the stroma. This mechanism allows the leukemic cells to evade systemic chemotherapy (Figure 1). Clinical data indicates that approximately 25 percent to 30 percent of AML patients exhibit abnormally elevated CXCR4 expression. This abnormally high expression can correlate with a significantly higher risk of disease relapse and broad chemoresistance.
Current treatments for AML?
Standard therapeutic intervention for Acute Myeloid Leukemia generally involves intense induction chemotherapy followed by consolidation therapy. Consolidation therapy frequently utilizes cytarabine to eliminate remaining leukemic blasts after initial remission is achieved. Other companies are looking at developing menin inhibitors because of their role in leukemogenesis. Motixafortide is another drug that has shown promising therapeutic effects for other hematological diseases like multiple myeloma that could be explored for AML treatment as well. However, despite these intense regimens, leukemic stem cells can still persist due to their protective anchoring within the bone marrow.
The use of motixafortide for AML treatment
To counteract the protective anchoring of AML cells, researchers have developed targeted CXCR4 inhibitors. This CXCR4 inhibitor was designed to "flush" leukemic stem cells out of their hiding spots to enhance chemotherapy response. Motixafortide is a selective CXCR4 inhibitor designed to block the interaction between the CXCR4 receptor and the CXCL12 ligand. By disrupting this critical biological axis, motixafortide mobilizes leukemic cells away from the protective bone marrow niche and forces a portion of leukemic cells into peripheral blood circulation. This mobilization is intended to disrupt the survival advantage these cells acquire, rendering them highly vulnerable to conventional chemotherapy like cytarabine. However, as we will see, patient stratification based on CXCR4 expression levels may be necessary to optimize the therapeutic benefit of motixafortide.
What are different endpoints evaluated for AML treatment
Clinical trials evaluating targeted treatments for AML utilize highly specific survival and molecular endpoints to measure therapeutic efficacy.
- RFS (Relapse-Free Survival): RFS serves as a primary endpoint, measuring the duration of time after a patient receives treatment for AML which they survive without any clinical signs of leukemic recurrence.
- OS (Overall Survival): Overall survival is a critical endpoint that calculates the total time from the date of diagnosis until death from any cause.
- MRD (Minimal Residual Disease): MRD quantification measures the minute fraction of leukemic cells remaining in the body post-treatment when the patient is in remission. MRD status could be utilized as a predictive marker for potential disease recurrence.
- CXCR4 as a biomarker: Evaluating the specific baseline and dynamic expression levels of CXCR4 protein on target cells serves as an exploratory biomarker to identify sub-populations that are most likely to respond positively to targeted blockade.
What is Minimal Residual Disease in AML, and Why is Accurate Detection So Difficult?
To understand the best methods to detect MRD for AML, one must examine the clinical landscape of remission and failure. Between 60 percent to 80 percent of all AML patients successfully achieve complete clinical remission following their initial induction therapy. Unfortunately, among the patients who achieve this complete remission, 50 percent to 70 percent will eventually experience disease relapse. These exceedingly high relapse rates emphasize the persistent biological nature of AML and highlight the severe limitations of current chemotherapeutic treatments.
Patients who test positive for Minimal Residual Disease (MRD) face a significantly higher clinical risk of relapse and demonstrate statistically shorter overall survival times compared to patients who are MRD negative. Even when MRD is classified as undetectable by standard clinical metrics, residual leukemic cells can still be present within the patient. Furthermore, residual pre-leukemic clones might persist and facilitate clonal hematopoiesis rather than full leukemia. Distinguishing between these cellular states remains a profound limitation for accurate diagnosis and sequential treatment.
Standard Methods for MRD Detection in AML and Their Clinical Limitations
Traditional analytical tools utilized for MRD detection carry distinct strengths but suffer from fundamental technical limitations that obscure total clonal architecture.
- Multiparameter flow cytometry (MFC): MFC is a widely utilized method for AML MRD assessment. However, this diagnostic tool frequently struggles to accurately distinguish between regenerating healthy precursor cells, normal clonal hematopoiesis, and residual leukemic blasts. Furthermore, MFC is largely unable to fully predict the true leukemic potential of phenotypically abnormal cellular populations.
- RT-qPCR: Real-time quantitative PCR delivers highly sensitive and highly specific molecular detection. The critical weakness of RT-qPCR is that it can only be effectively applied to a subset of AML patients with trackable molecular markers, detectable leukemia-specific mutations or genetic fusions.
- Bulk NGS: Next-generation sequencing enables simultaneous detection of multiple genetic mutations across large gene panels, providing a comprehensive view of the mutational landscape within a sample. However, because bulk NGS measures averaged signals across thousands to millions of cells, it cannot determine which mutations co-occur within the same individual cells or definitively resolve clonal architecture. While bulk NGS can detect persistent mutations during remission, it cannot distinguish whether these mutations reside in true leukemic cells, pre-leukemic clones, or non-malignant clonal hematopoiesis without single-cell resolution.
Why Combine Single-Cell DNA Sequencing and Immunophenotyping for AML MRD Detection?
To circumvent the inherent limitations of the standard methods mentioned above, an emerging approach with growing translational and clinical utility applications is single-cell multiomics. Using a multiomic workflow that analyzes DNA and protein from the same cells helps researchers distinguish between true leukemic cells, pre-leukemic clones, and those that are wildtype. Some platforms, like Mission Bio’s Tapestri® platform, even have the capability to simultaneously process both DNA sequence data and targeted protein expression within each individual cell giving current standard of care methods a meaningful advantage over conventional approaches.
Why Target the CXCR4 and CXCL12 Axis for AML MRD Detection in the BLAST Trial?
Data presented at the European Hematology Association (EHA) Congress 2025 from the Heidelberg University study on the SAL BLAST trial was also highlighted in a webinar demonstrating the power of Mission Bio’s single-cell DNA + Protein platform providing retrospective clinical trial insights suggesting CXCR4 expression may serve as a predictive biomarker for AML patient stratification. The BLAST study was structured as a randomized, double-blind, placebo-controlled phase II clinical trial between 2015 and 2019 at 29 sites in Germany consisting of 128 patients. During enrollment, these patients were randomized to receive either Motixafortide or a placebo alongside consolidation therapy, cytarabine (figure 2).
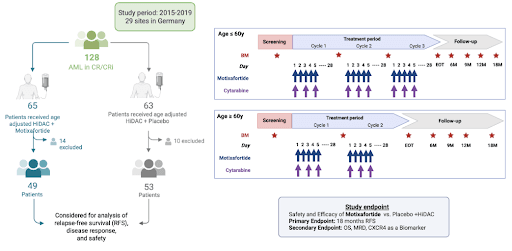
Of note, patients included in the trial had to be in their first clinical remission following initial induction chemotherapy. The patient cohort was carefully stratified by age. Patients aged below 60 received up to three full cycles of chemotherapy, whereas patients aged over 60 received two cycles. During the trial protocols, Motixafortide was administered subcutaneously precisely before the delivery of cytarabine.
Researchers recognized that testing entire cohorts using generalized metrics could conceal specific biological responses. They specifically focused on the CXCR4/CXCL12 axis because of its proven function in facilitating cell retention within the protective bone marrow niche. The researchers aimed to deeply investigate the relationship between localized CXCR4 activity and the emergence of MRD subclones, focusing on discovering the precise mechanisms of molecular treatment resistance.
Why Did the Initial BLAST Trial Data Fail to Show a Survival Benefit in AML Patients?
Initial clinical evaluations of the bulk trial data yielded highly disappointing outcomes: when researchers looked at the overall patient population, there was no statistical difference in relapse-free survival between the drug and the placebo. Deeper analysis, however, revealed important insights into why the unselected cohort showed no benefit.
No improvement with CXCR4 inhibition in unselected AML patient cohorts
When the clinical team evaluated the entire patient cohort, the primary trial endpoint of relapse-free survival failed to separate the treatment arms (figure 3). The median relapse-free survival duration was measured at 10.3 months for the motixafortide group and 11.5 months for the placebo control group. The overall survival metrics also demonstrated no significant statistical enhancement with the implementation of the CXCR4 inhibitor.
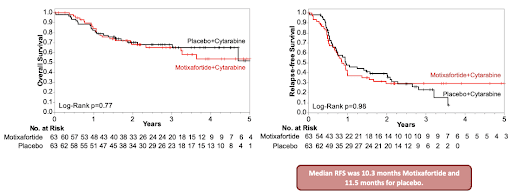
Acute myeloid leukemia is not one disease per patient but a diverse clonal ecosystem
Single-cell analysis revealed that AML operates as a highly complex biological ecosystem. Minor subclones that represent a tiny fraction of cells during remission can eventually dominate, causing relapse. The single-cell multiomics platform revealed vast cellular heterogeneity across the AML samples, mapping T-cells, NK-cells, monocytes, and the highly critical blast compartment (Figure 4). This allowed the scientists to see which cell contains which mutation at the designated timepoint, which is advantageous when understanding how clonal evolution occurs in the complex diseases such as AML.
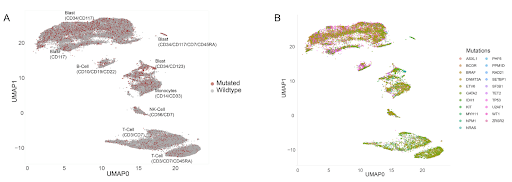
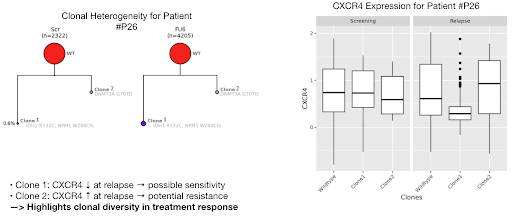
By generating intricate phylogenetic trees for individual patients, researchers mapped exact clonal architecture as depicted in Figure 5 on the left for patient #P26. Subclones in a variety of patients carrying mutations such as DNMT3A, TET2, and PPM1D expanded drastically by the time of clinical relapse. Furthermore, different clones within the same patient reacted differently to motixafortide, where one clone might decrease its CXCR4 expression while another might increase it, suggesting differential clone-specific biological responses to treatment as depicted in Figure 5 on the right. This highlights that selective therapeutic pressures dynamically reshape signaling pathways resulting in changes in the survival of distinct subclones.
In the case of patient #P26, single-cell multiomic analysis permitted researchers to quantify MRD reliably and observe the real-time evolutionary tracks of specific mutated clones. Clone 1 featured a documented decrease in CXCR4 protein expression at the time of relapse. This decrease suggests a degree of biological sensitivity to the motixafortide intervention. Concurrently, Clone 2 within the exact same patient demonstrated an increase in CXCR4 expression at the point of relapse. This isolated increase implies severe localized cellular resistance to the therapy. This clone-specific tracking confirms that single-cell MRD approaches can precisely characterize the genotypic and phenotypic features of residual disease.
CXCR4 expression serves as a predictive biomarker to stratify patients via single-cell analysis
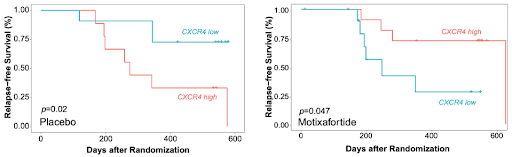
When patients were retrospectively categorized based strictly on their cellular CXCR4 expression levels, clinical outcomes diverged dramatically. Patients with high CXCR4 expression who received motixafortide demonstrated a significantly improved relapse-free survival rate. Conversely, the drug provided no benefit to patients with low CXCR4 expression, whose outcomes were actually worse than the placebo control group (Figure 6). The interaction term examining the relationship between the treatment arm and specific CXCR4 status yielded a highly significant p-value of 0.001. These results suggest potential clinical utility validation of single-cell multiomics for AML patient stratification. By simultaneously evaluating both genotypic and phenotypic profiles at the single-cell level, researchers can tailor targeted therapies to specific biological signatures rather than using a one-size-fits-all approach for everyone with the same disease (Figure 6).
Clinical utility implications for single-cell multiomics and MRD detection in acute myeloid leukemia
The results derived from the BLAST trial reanalysis carry extensive implications for the immediate future of hematology and oncology. The results suggest that future use of motixafortide should focus on CXCR4 expression as a predictive biomarker and should be reserved for patients whose leukemia expresses high levels of CXCR4. By abandoning traditional bulk processing metrics, scientists can prevent the statistical masking of highly effective targeted therapies.
Single-cell multiomics directly addresses the limitations of standard flow cytometry and bulk sequencing by combining deep genotypic profiling with precise surface protein quantification. This simultaneous measurement greatly improves the ability to distinguish between leukemic, pre-leukemic, and non-malignant clonal populations. Moving forward, clinical trial designs should incorporate single-cell MRD assessment not merely as an exploratory endpoint, but as a foundational metric for patient stratification.
Beyond the scope of the SAL BLAST trial, this data proves that high-resolution single-cell analysis is essential for identifying which specific patients will actually benefit from targeted therapies. By identifying the exact leukemic subclones that will respond to specific interventions, scientists can help guide more precise deployment of targeted therapies. Implementing this high-resolution single-cell MRD assessment for diseases like AML not only maximizes therapeutic efficacy but also spares patients from the unnecessary toxicity of ineffective treatments, establishing a more efficient standard of care.
Contact us today to discover how single-cell multiomics can uncover hidden biomarkers, improve patient stratification, and deliver true clinical utility for your MRD research.
Explore acute myeloid leukemia products and services:
Acute Myeloid Leukemia (AML) DNA Panel
Targeted Single-cell RNA Panel
TotalSeq™-D Acute Myeloid Leukemia Antibody Cocktail
Custom, fit-for-purpose assays






